Development of a prediction model for student teaching satisfaction based on 10 machine learning algorithms
Statistical methods
SHAP technology
SHAP is a model interpretation technique based on game theory that reveals the contribution of each feature to the model’s prediction results, aiming to distribute the contribution of features to the prediction results fairly. Its core idea is to treat each feature as a “participant, ” calculate the marginal contribution of all possible feature combinations to the prediction results, and then obtain the SHAP value of each feature through weighted averaging. This method combines global explanations (such as feature importance rankings) with local explanations (such as the basis for predicting individual samples), providing multi-dimensional model transparency.
SHAP can elucidate nonlinear relationships and interactions among features. Being model-agnostic, SHAP can be utilized with any predictive model. It employs Shapley values to articulate the contribution of each input feature to the predictions made by the model. The Shapley value, defined as the average marginal contribution of feature values across all conceivable coalitions, is computed as demonstrated in Eq. (1).
$$\phi i(f,x)=\sum\nolimits_{{z^{\prime} \subseteq x^{\prime}}} {\frac{{\left| {z^{\prime}} \right|!(M – \left| {z^{\prime}} \right| – 1)!}}{{M!}}\left[ {{f_x}(z^{\prime}) – {f_x}(z^{\prime}\backslash \{ i\} )} \right]}$$
(1)
\(\phi i(f,x)\) represents the Shapley value contribution of feature i to the prediction result of model f for a specific input x. \(z^{\prime}\) is a subset of the input feature set \(x^{\prime}\), which \(x^{\prime}\) generally consists of features other than target feature i. M represents the total number of features.
\({f_x}(z^{\prime})\) refers to the model’s predicted values under a feature subset \(z^{\prime}\), where the feature subset \(z^{\prime}\) is selected from the original input x. \({f_x}(z^{\prime}\backslash \{ i\} )\) is the predicted value of the model under a subset that does not include feature i. \(\frac{{\left| {z^{\prime}} \right|!(M – \left| {z^{\prime}} \right| – 1)!}}{{M!}}\) is the weight of subset \(z^{\prime}\), which reflects the proportion of importance of this subset among all possible combinations.\({f_x}(z^{\prime}) – {f_x}(z^{\prime}\backslash \{ i\} )\) is the marginal contribution to the predicted value when the x feature is added to subset \(z^{\prime}\).
K-means algorithm
K-means is an unsupervised learning algorithm that aims to divide n data points into K mutually exclusive clusters so that data points within each cluster are as close as possible (high intra-cluster similarity) and clusters are as dispersed as possible (low inter-cluster similarity). Its core idea is to minimize the within-cluster sum of squares (WCSS) through iterative optimization, which is the sum of the squares of the distances between data points and their respective cluster centers.
Let the dataset is\(X=\{ x1,x2,….,xn\}\), each data point \(xi \in {{\mathbb{R}}^d}\)(d is the feature dimension), and the cluster center set be \(C=\{ c1,c2,…,cK\}\), where \({c_k} \in {\mathbb{R}}{\ominus ^d}\) is the center of the k cluster.
The distance between data points \(xi\)and cluster centers \({c_k}\) is usually measured using Euclidean distance, which is calculated using the formula in Eq. (2).
$$d({x_i},{c_k})=\sqrt {{{\sum\nolimits_{{j=1}}^{d} {({x_{ij}} – {c_{kj}})} }^2}}$$
(2)
Among them, \({x_{ij}}\) is the j feature value of the data point\(xi\), and \({c_{kj}}\)is the j feature value of the cluster center\({c_k}\). The algorithm iterates repeatedly, assigning each data point \(xi\)to the cluster corresponding to the nearest cluster center \({c_k}\)and updating the cluster centers until a stopping condition is met, such as the change in cluster centers being less than a certain threshold or reaching a preset number of iterations.
Boruta algorithm
The Boruta algorithm is a feature selection algorithm designed to select the most important features for the prediction target from high-dimensional data sets. Its core idea is to screen features by comparing the importance of the original features with randomly generated “shadow features” in the random forest model.
Assuming we have T decision trees, the formula \(I(X)\)for calculating the importance of feature X can be expressed as Eq. (3):
$$I(X)=\frac{1}{T}\sum\nolimits_{{t=1}}^{T} \Delta {G_t}(X)$$
(3)
Among them, \(\Delta {G_t}(X)\)is the reduction in the Gini index caused by splitting using feature X in the t decision tree.
Suppose there are n importance values \({I_1},{I_2},{I_3},…,{I_n}\) for the original features and m importance values \({J_1},{J_2},{J_3},…,{J_m}\) for the shadow features. We need to test whether the mean importance value of the original features\({\mu _I}\) is greater than the mean importance value of the shadow features\({\mu _J}\). The formula for the t-test is given by Eq. (4):
$$t=\frac{{I – J}}{{{S_p}\sqrt {\frac{1}{n}+\frac{1}{m}} }}$$
(4)
Among them, \(I=\frac{1}{n}\sum\nolimits_{{i=1}}^{n} {{I_i}}\), \(J=\frac{1}{m}\sum\nolimits_{{i=1}}^{m} {{J_j}}\), \({S_p}\) are the combined standard deviations, the calculation formula is given by Eq. (5).
$${S_p}=\sqrt {\frac{{(n – 1)s_{I}^{2}+(m – 1)s_{J}^{2}}}{{n+m – 2}}}$$
(5)
where\(s_{I}^{2}\)and\(s_{J}^{2}\) are the sample variances of the importance values of the original features and shadow features, respectively. The calculated t-value is compared with the critical value at a given significance level. If t is greater than the critical value, the importance of the original feature is considered to be significantly higher than that of the shadow feature.
The algorithm determines important features through multiple iterations. This process continues until all features are classified as important or unimportant, or until the preset number of iterations is reached.
Dataset and feature selection
The dataset used in this paper is sourced from the Turkish Student Evaluation Project28. It includes multidimensional features such as students’ classroom participation, homework completion, and exam scores. The dataset contains 5,820 records, each comprising 33 feature variables.
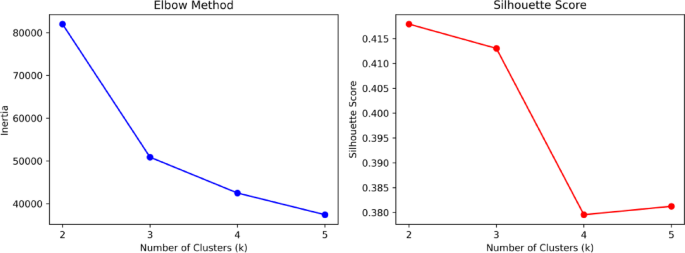
The dataset itself is of good quality with no missing values. To investigate the impact of each metric on student evaluation outcomes, we applied the K-means clustering algorithm to the dataset. Figure 1 shows the elbow plot and silhouette coefficient plot of the dataset. Based on the optimization results, the dataset is divided into three categories. The distribution plot of the number of each category is presented in Fig. 2, with the counts of each category being 1232, 2230, and 2358 respectively. The number distribution across the categories is relatively balanced.
Using the Boruta algorithm, we selected 30 essential features from the 34 fields, including features Q10, and Q23, for subsequent machine learning. After applying Boruta, we found that feature Q10 consistently showed higher importance scores than its shadow features across multiple runs. Compared to other features, feature Q23 made a greater contribution to reducing prediction errors in the model. The other selected features also conformed to the screening rules of the Boruta algorithm. Figure 3 shows the weight ranking information of the 30 features selected by the Boruta algorithm, where a higher ranking indicates a greater influence of the weight.
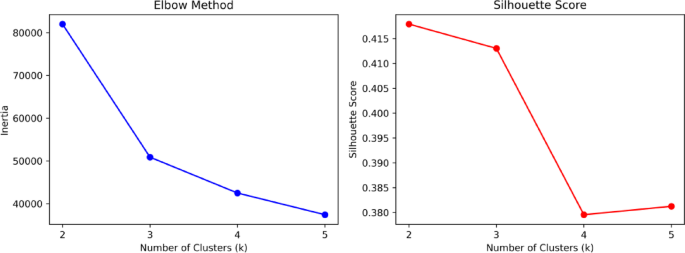
Elbow diagram and contour coefficient diagram.
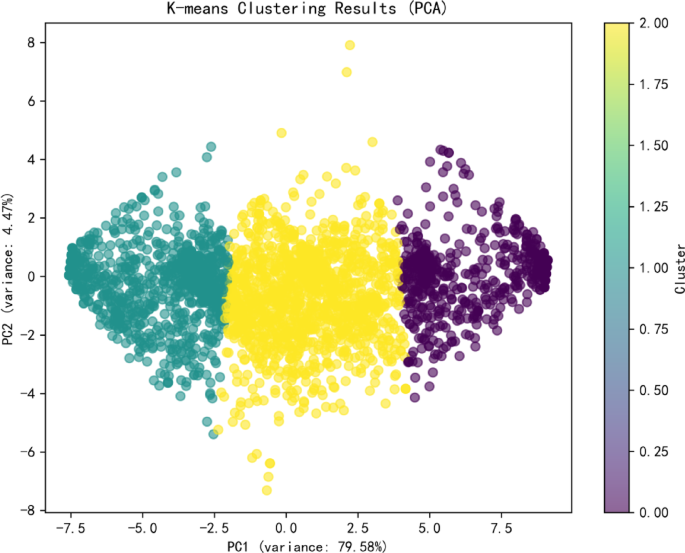
K-means clustering diagram.
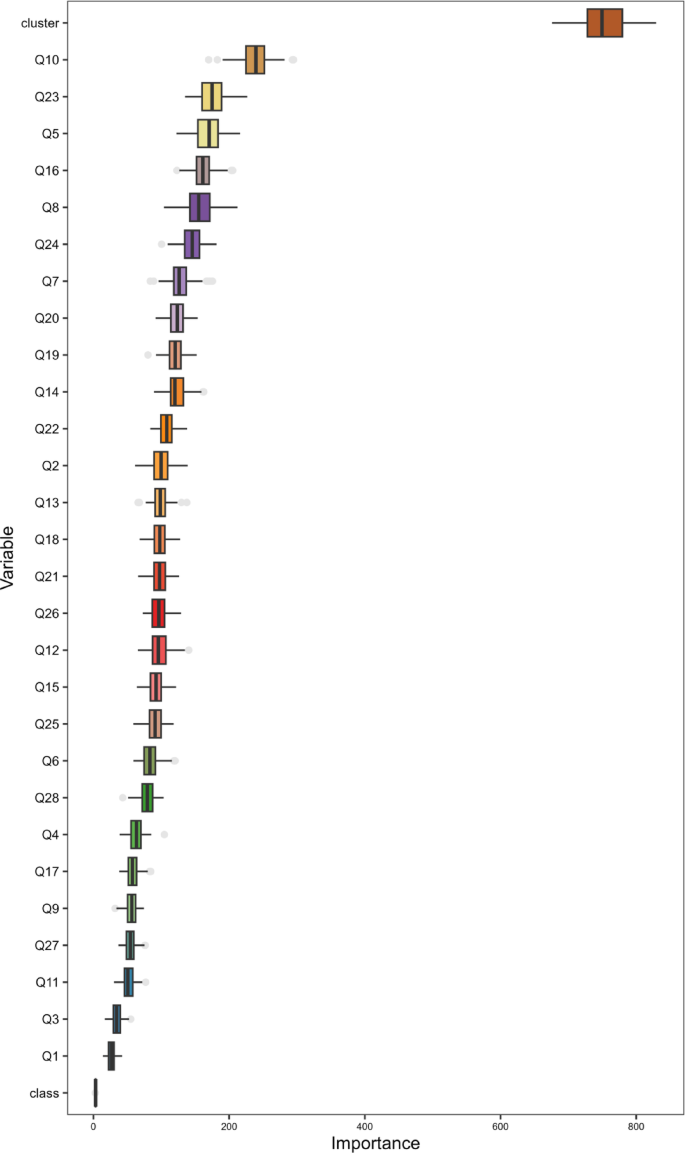
Evaluation indicators selected based on the Boruta algorithm.
Technical approach
For this academic performance prediction task, we use a dataset containing 5820 sample. The dataset is divided into training and testing sets using the random function sample, with 70% allocated to the training set and 30% to the testing set, which can strike a balance between “model learning sufficiency” and “generalization evaluation reliability”. Meanwhile, the subsequent use of 5-fold cross-validation for further model optimization is also compatible with this 70 − 30 split.
The study employs 10 machine learning algorithms, such as RF, GBM, NB, KNN, Nnet, FDA, SVM, CART, SLDA, and ADA, to predict and evaluate student learning outcomes. Each model undergoes 5-fold cross-validation or parameter optimization (For the random forest algorithm, the hyperparameters in the grid search were configured as mtry is (2, 5, 10), splitrule is gini, min.node.size is (1, 5, 10). For the SVM algorithm, the optimal cost and gamma parameters are searched through 5-fold cross-validation. The candidate values for the cost parameter are (0.1, 1, 10, 100, 1000), and the candidate values for the gamma parameter are (0.001, 0.01, 0.1, 1, 10).
We evaluate the model’s performance on the test set using machine learning evaluation metrics such as Accuracy (95% CI), Sensitivity, Specificity, PPV, NPV, Precision, Recall, and F1.
All analyses in this paper primarily use R software (4.3.0) and Python software (3.11.0). We utilized packages such as plotROC, caret, pROC, and e1071 in R software to plot ROC curves, and used Python software to plot SHAP plots for parameter visualization. Finally, we employed the shiny package to develop an online learning effectiveness satisfaction prediction app.
Figure 4 presents the technical workflow of this study. The dataset is partitioned. 10 machine learning algorithms are adopted for comparative analysis. Finally, feature visualization is performed using SHAP and Shiny.
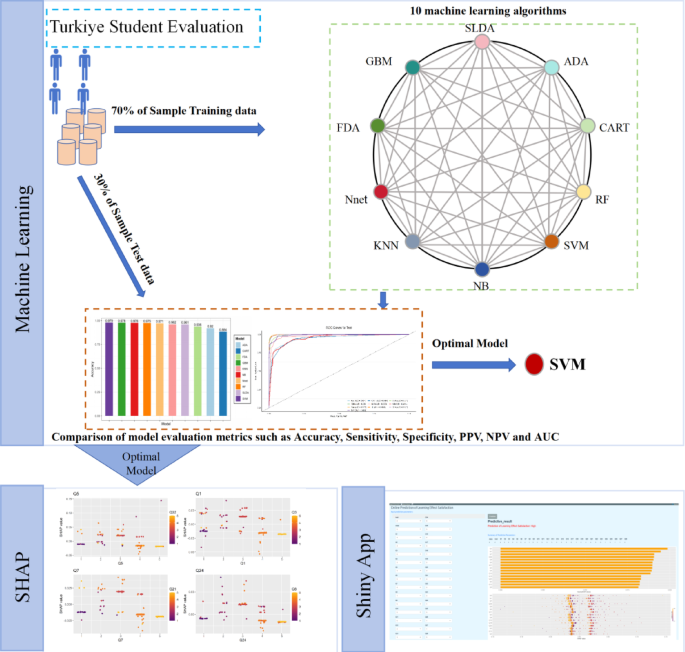
Research methodology diagram.
Evaluation metrics
To assess the efficacy of the prediction model, we employ standard metrics frequently utilized in student grade prediction frameworks, specifically accuracy, precision, recall, and F1 score29.
True Positive (TP) refers to the count of genuinely positive instances that the model accurately identifies as positive.
False Positive (FP) denotes the number of instances that are actually negative but are incorrectly classified as positive by the model.
True Negative (TN) indicates the number of genuinely negative instances that are correctly identified as negative by the model.
False Negative (FN) represents the count of positive instances that are misclassified as negative by the model.
Precision is shown in Eq. (6):
$$Precision=\frac{{TP}}{{TP+FP}}$$
(6)
The proportion of truly positive samples among those predicted to be positive reflects the accuracy of the model’s positive predictions.
The recall rate formula is shown in Eq. (7):
$$Recall=\frac{{TP}}{{TP+FN}}$$
(7)
The proportion of samples that are actually positive examples and are correctly predicted as positive examples by the model measures the model’s ability to capture positive examples.
Specificity is shown in Eq. (8):
$$Specificity=\frac{{TN}}{{TN+FP}}$$
(8)
The proportion of samples that are actually negative examples and are correctly predicted as negative examples by the model reflects the model’s ability to identify negative examples.
The accuracy formula is shown in Eq. (9):
$$Accuracy=\frac{{TP+TN}}{{TP+TN+FP+FN}}$$
(9)
The proportion of samples correctly predicted by the model out of the total number of samples reflects the model’s overall predictive accuracy.
The F1 score formula is shown in Eq. (10):
$$F1=2 \times \frac{{Precision \times Recall}}{{Precision+Recall}}$$
(10)
F1 combines precision and recall metrics to balance their importance, with values ranging from 0 to 1. The closer the value is to 1, the better the model performance.
link







